In-silico Assessment of Arrhythmogenic Potential of Atrial Ablation Patterns
- Ansprechperson:
- Projektgruppe:
Herzmodellierung
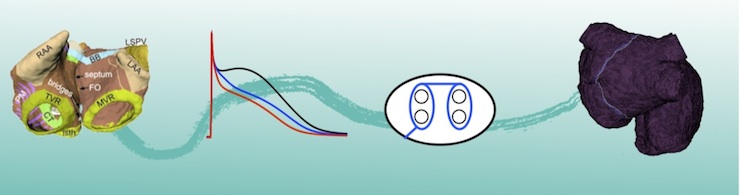
Atrial fibrillation (AF) and Atrial flutter (AFlut) are the most common cardiac arrhythmias affecting more than 1% of the population. Even though they are not immediately life threatening, they are associated with severe complications such as cerebral stroke. Radio frequency or cryo-ablation are the only curative therapies for these diseases. However, many patients who underwent AF ablation develop AFlut subsequently. The aim of this work is to predict the changes in vulnerability to AFlut induced by certain ablation scar patterns in-silico.
Towards this end, personalized models are built based on patient-specific cardiac imaging data and enhanced with rule-based fiber orientation and tissue labels. Further individualization can include the patient’s AF history (degree of AF-induced remodeling), medication, and genetic information.
The arrhythmogenic potential of certain ablation patterns can then be assessed by dedicated algorithms using the patient-specific model.
Publications
- M.W. Krueger et al., Personalization of Atrial Anatomy and Elelectophysiology as a Basis for Clinical Modeling of Radio-Frequency-Ablation of Atrial Fibrillation. IEEE Transactions on Medical Imaging (31), 73-84, 2012
- A. Loewe et al., Arrhythmic Potency of Human Ether-à-go-go-related Gene Mutations L532P and N588K in a Computational Model of Human Atrial Myocytes. Europace (16), 435-443, 2014
- G Seemann et al., Atrial Fibrillation-based Electrical Remodeling in a Computer Model of the Human Atrium. Proceedings Computing in Cardiology (37), 417-420, 2010
- M.W. Krueger et al., In-silico Modeling of Atrial Repolarization in Normal and Atrial Fibrillation Remodeled state. Medical & Biological Engineering & Computing (51), 1105-1119, 2013

