Methods for the Integration of Ionic Current Measurement Data into Models of Cardiac Electrophysiology
- type:Bachelor thesis
- tutor:
- person in charge:
-
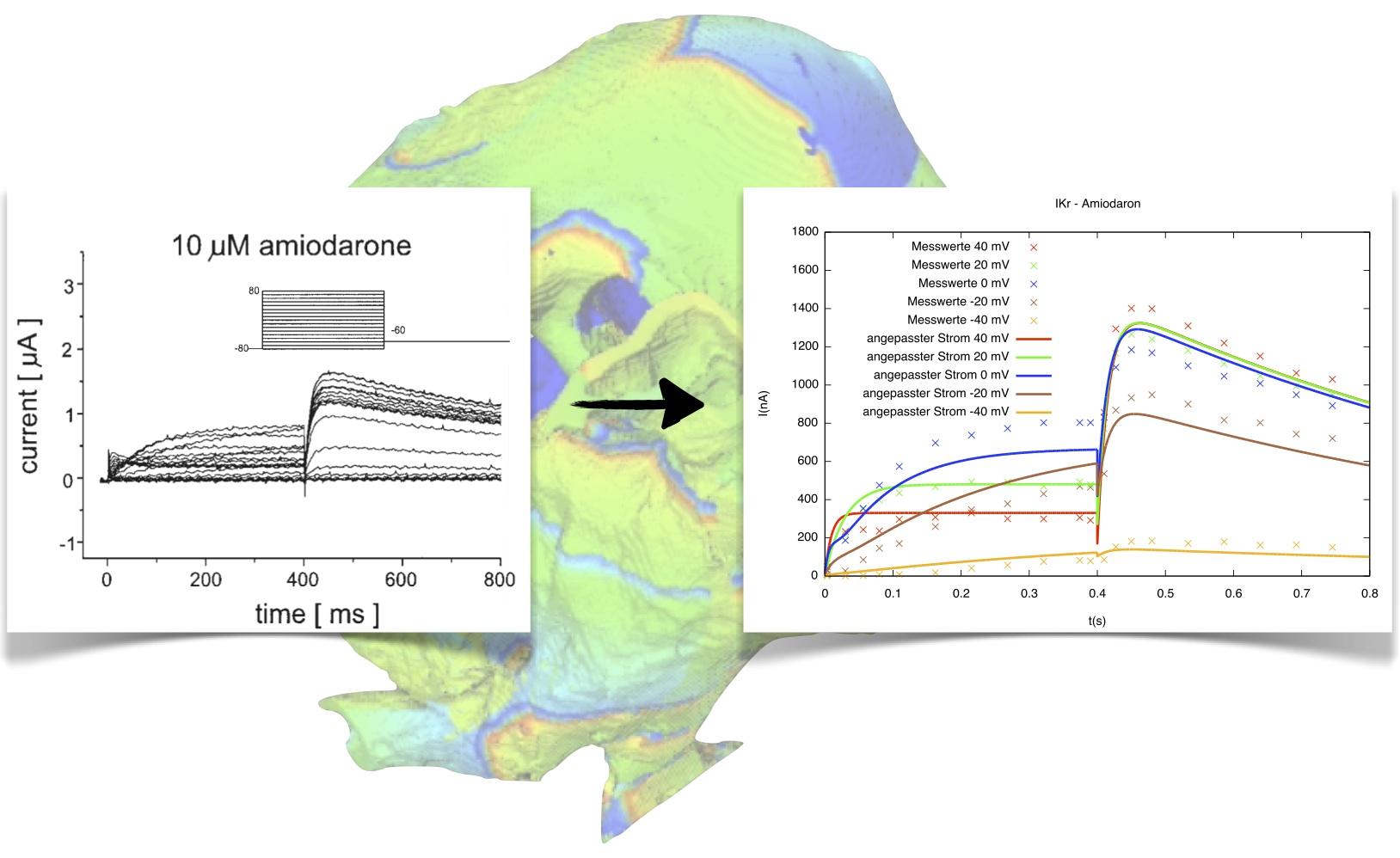
Most of the current models of cardiac electrophysiology are based on the work of Hodgkin and Huxley (1952), where a mathematical description of ion currents of the membrane of a giant squid axon is given. More sophisticated models include descriptions of ion channels using Markov chains. Generally, these published models reflect the behavior of only one certain cell type and species under physiologic conditions. However, the focus of research is, of course, on unphysiologic conditions, such as mutations of ion channels, or under the influence of drugs. As a consequence, the electrophysiological models have to be modified to reflect these other conditions. For this purpose, the parameters of the ion channels have to be adjusted, so that the simulated ion currents are fitted to the measured unphysiologic ones.
In previous work, a modular framework for any parameter fitting was developed at the Institute of Biomedical Engineering. Furthermore, a MATLAB environment for the adjustment of certain ion channels was drafted. However, the optimization process of cardiac ion channels can be very time-consuming depending on the number of parameters. Therefore, the MATLAB optimization has to be improved and compared to the modular parameter fitting framework. Finally, different measurement protocols should be investigated, compared and evaluated regarding the efficiency of integrating data into the model.